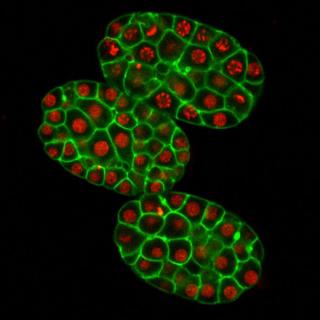
The BMSE program encompasses a wide variety of research within the broad areas of molecular genetics and molecular biology. Specific areas of research include : (1) the ecology , regulation, and systems biology of diseases and symbioses caused by viruses, bacteria, and fungi in plants and animals and (2) understanding developmental processes, regulatory networks, ion channels, microtubules, neural plasticity, and stem cell biology in invertebrates such as nematodes and vertebrates including marine chordates and mammals.





















